Please cite:
Schudoma et al.,
Nucl. Acids Res. 38: 970-980.
DOI 10.1093/nar/gkp1010.
Schudoma et al.,
Nucl. Acids Res. 38: 970-980.
DOI 10.1093/nar/gkp1010.
| '2-2-Internal Loop pdb1xnq1A.n1386-1453 | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Source: [PDB-id:chain] | 1xnq:A (&rarr PDB) | ||||||||||||||||||||||||||||||
| Source: Information | STRUCTURE OF AN INOSINE-ADENINE WOBBLE BASE PAIR COMPLEX IN THE CONTEXT OF THE DECODING CENTER | ||||||||||||||||||||||||||||||
| Source: Compound | 16S RIBOSOMAL RNA | ||||||||||||||||||||||||||||||
| Source: Resolution | 3.05 ANGSTROMS. | ||||||||||||||||||||||||||||||
| Position | (1388, 1451), (1391, 1448) | ||||||||||||||||||||||||||||||
| Primary structure ('_': anchors) | _GA_-_GA_ | ||||||||||||||||||||||||||||||
|
Bases with unusual sugar puckers (Standard: C3'-endo) |
None | ||||||||||||||||||||||||||||||
|
Bases with unusual glycosidic-bond configuration (Standard: anti) |
None | ||||||||||||||||||||||||||||||
| Tertiary structure: Stacked bases |
|
||||||||||||||||||||||||||||||
|
Tertiary structure: Base-pairs (anchor pairs) |
|
||||||||||||||||||||||||||||||
| Downloads |
Atom coordinates (PDB format) Contact annotation (MC-Annotate format) |
||||||||||||||||||||||||||||||
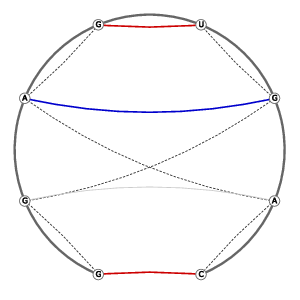
| 3D Structure | Structure Graph | ||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||
| Structural Clusters | |||||||||||||||||||||||||||||||


