Please cite:
Schudoma et al.,
Nucl. Acids Res. 38: 970-980.
DOI 10.1093/nar/gkp1010.
Schudoma et al.,
Nucl. Acids Res. 38: 970-980.
DOI 10.1093/nar/gkp1010.
| '4-2-2-0-5-Multiloop pdb1njm10.m538-1208 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Source: [PDB-id:chain] | 1njm:0 (&rarr PDB) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Source: Information | THE CRYSTAL STRUCTURE OF THE 50S LARGE RIBOSOMAL SUBUNIT FROM DEINOCOCCUS RADIODURANS COMPLEXED WITH A TRNA ACCEPTOR STEM MIMIC (ASM) AND THE ANTIBIOTIC SPARSOMYCIN | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Source: Compound | 23S RIBOSOMAL RNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Source: Resolution | 3.60 ANGSTROMS. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Position | (543, 1203), (548, 631), (634, 772), (775, 1143), (1144, 1197) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Primary structure ('_': anchors) | _GACU_-_GA_-_UU_-__-_CGGAU_ | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Bases with unusual sugar puckers (Standard: C3'-endo) |
None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Bases with unusual glycosidic-bond configuration (Standard: anti) |
None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tertiary structure: Stacked bases |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Tertiary structure: Base-pairs (anchor pairs) |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Downloads |
Atom coordinates (PDB format) Contact annotation (MC-Annotate format) |
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3D Structure | Structure Graph | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
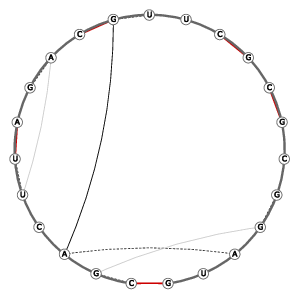
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structural Clusters | |||||||||||||||||||||||||||||||||||||||||||||||||||||||


